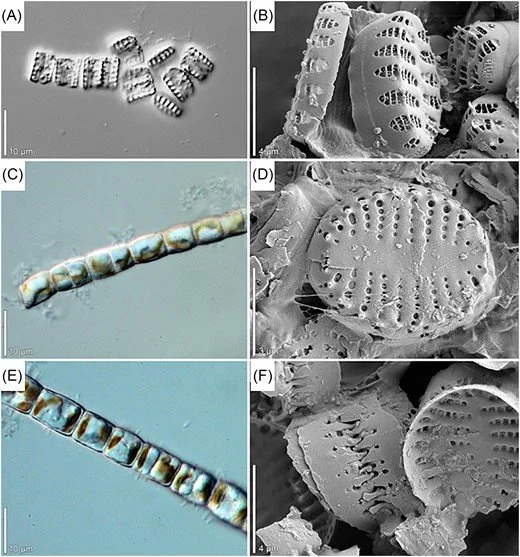
Figure source: Mazur-Marzec, H., Andersson, A. F., Błaszczyk, A., Dąbek, P., Górecka, E., Grabski, M., ... & Węgrzyn, A. (2024). Biodiversity of microorganisms in the Baltic Sea: the power of novel methods in the identification of marine microbes. FEMS microbiology reviews, 48(5), fuae024.
Analyzing Microbiome Metabarcoding Data with TabPFN
TabPFN [1] is a powerful, novel foundation model for tabular data. It it is currently revolutionizing ML for small, tabular data, which was previously dominated by classical methods like XGBoost. We would like to investigate the abilities of TabPFN to model metabarcoding data [2] (species composition of biological communities, obtained by DNA sequencing analysis of environmental samples). Specifically, we’ll investigate TabPFNs ability to predict environmental variables from species composition, inspect the model’s latent representation and try multi-task learning.
Key Publications
[1] Hollmann, N., Müller, S., Purucker, L., Krishnakumar, A., Körfer, M., Hoo, S. B., ... & Hutter, F. (2025). Accurate predictions on small data with a tabular foundation model. Nature, 637(8045), 319-326.
[2] Zschaubitz, E., Schröder, H., Glackin, C. C., Vogel, L., Labrenz, M., & Sperlea, T. (2025). A benchmark analysis of feature selection and machine learning methods for environmental metabarcoding datasets. Computational and Structural Biotechnology Journal, 27, 1636-1647.
Contact
Stefan Lüdtke
stefan.luedtke@uni-rostock.de